Plasmid purification is often assessed using simple spectrophotometric metrics such as the A260/280 ratio, which primarily reflects protein contamination. However, this measurement provides only a narrow view of purity. In practice, high quality plasmid DNA must be free not only of proteins, but also of endotoxins, genomic DNA, RNA, and residual salts. These impurities can remain undetected during basic quality checks while still exerting significant influence on downstream applications.
A more complete definition of clean plasmid DNA encompasses the removal of all components capable of altering enzymatic reactions, affecting cell health, or introducing variability into experimental workflows.
How impurities influence qPCR, NGS, and transfection workflows
Downstream applications are highly sensitive to contaminants that may not be apparent during routine quantification.
In mammalian cell transfection, endotoxins are a particularly important consideration. Even at low levels, they can reduce cell viability and interfere with uptake of genetic material, leading to inconsistent expression outcomes despite apparently adequate plasmid yield.
For qPCR and NGS, contaminants such as salts, chaotropic agents, RNA, and genomic DNA can alter reaction efficiency or contribute to biased library preparation. These subtle effects often manifest as reduced amplification performance, increased variability across replicates, or diminished sequencing quality.
When plasmid DNA does not meet the necessary purity standards, the resulting downstream variability can obscure the true performance of reagents, constructs, or protocols.
The operational cost of inadequate plasmid purity
Insufficient plasmid purity frequently leads to repeated transfections, additional optimisation cycles, and unnecessary troubleshooting. These activities consume time, reagents, and personnel effort that could otherwise be directed to productive experimental work. In many cases, the underlying issue originates from upstream DNA quality rather than downstream workflows, but this connection may only become apparent after multiple unsuccessful attempts.
Ensuring high quality plasmid DNA from the outset is therefore an essential component of efficient laboratory operation and reliable biological interpretation.
How ZymoPURE™ II supports consistency and reproducibility
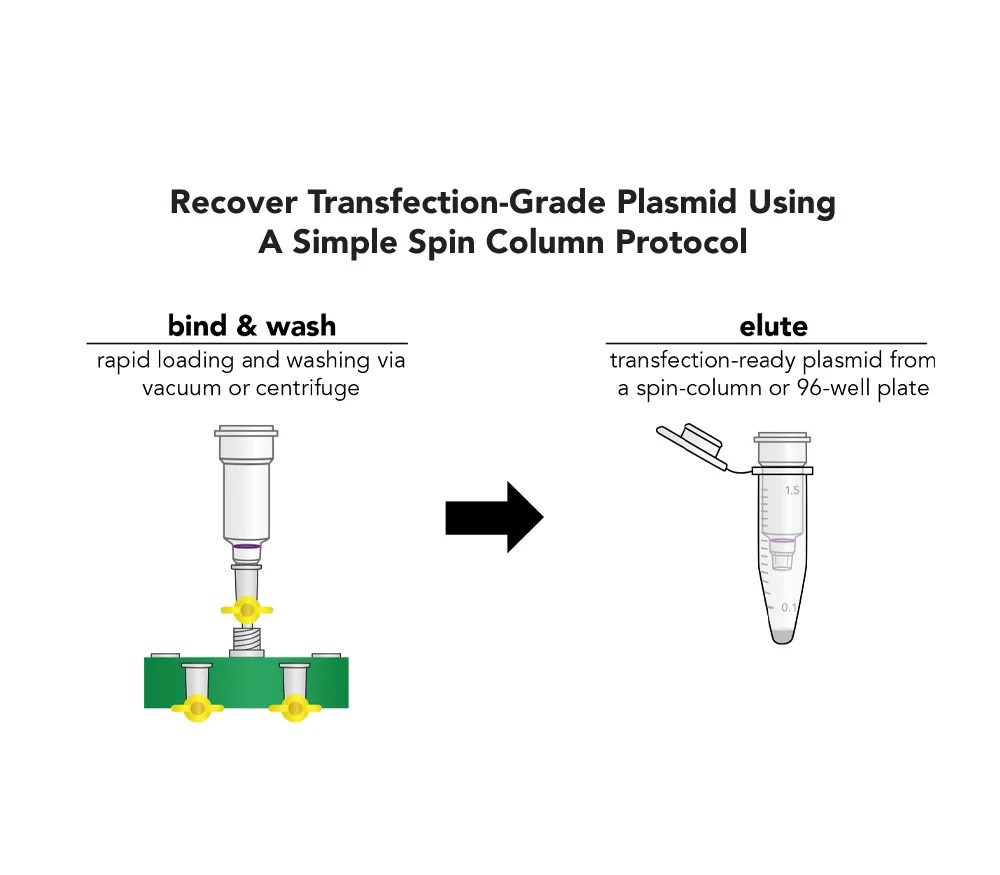
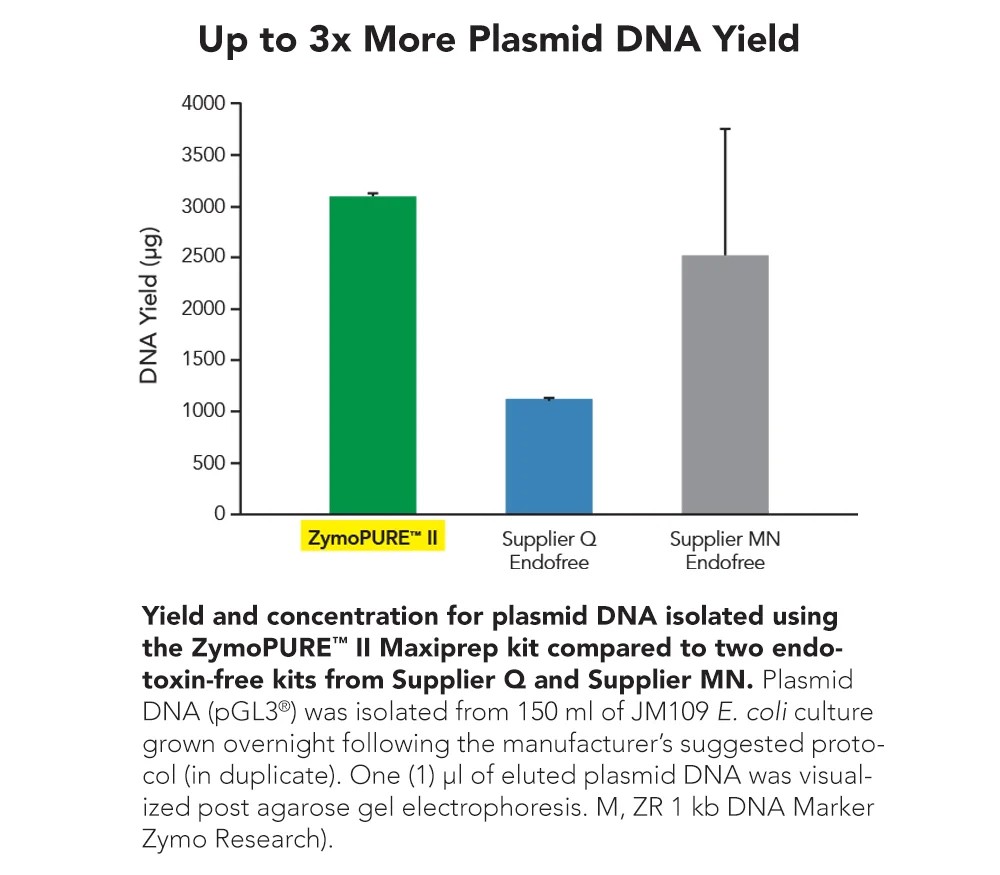
Zymo Research’s ZymoPURE II plasmid purification kits are designed to address these challenges directly. The kits employ a silica based purification system combined with an integrated endotoxin reducing workflow. This approach enables the removal of impurities that can interfere with biological assays, resulting in plasmid DNA that is suitable for demanding applications such as transfection, qPCR, and next generation sequencing.
|
|
|
|---|
A key advantage of the system is its reproducibility. The workflow produces consistent plasmid purity and performance across users, batches, and laboratories, reducing the variability that often arises from manual purification steps or incomplete contaminant removal.
The link between plasmid purity and experimental outcomes
High quality plasmid DNA provides a stable foundation for downstream processes. When contaminants are effectively removed, expression levels become more predictable, transfections yield more uniform results, and sequencing or qPCR workflows perform with greater reliability. This clarity enables researchers to draw more confident conclusions from their data and reduces the need for iterative troubleshooting.
Clean input directly supports clean biological outcomes. By adopting purification methods that remove all relevant impurities, researchers can ensure that plasmid quality supports rather than limits the performance of their experiments.
Explore how ZymoPURE kits can support your research, or contact our experts to with any queries.