Modern microbiome research depends heavily on Next-Generation Sequencing (NGS). These methods allow scientists to characterise complex microbial communities in remarkable detail. Despite their power, microbiome analysis pipelines are vulnerable to multiple sources of technical bias that can compromise both accuracy and reproducibility. One of the most impactful and frequently overlooked sources of error occurs during DNA extraction, specifically through a phenomenon known as lysis bias.
The challenge of lysis bias in DNA extraction
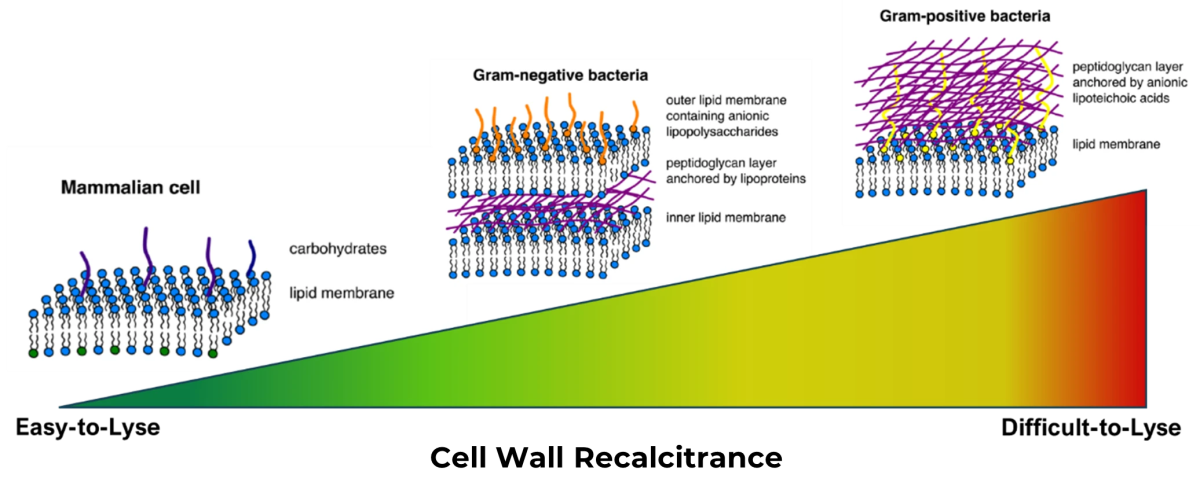
Lysis bias arises when different microorganisms are broken open, or lysed, with unequal efficiency, causing certain taxa to be under- or overrepresented in the resulting data. The ease with which a cell releases its DNA is dictated by the complexity of its cell envelope. Mammalian cells possess only a lipid membrane and are therefore relatively easy to disrupt. In contrast, Gram-positive bacteria have thick, highly structured peptidoglycan cell walls that require much more aggressive treatment to lyse effectively. At Zymo Research, this resistance to disruption is described as “cell wall recalcitrance,” a term used to categorise microorganisms by how difficult they are to break open. As cell wall recalcitrance increases, the likelihood of incomplete lysis, and therefore biased results, also increases.
Real-world evidence of bias in microbiome analysis
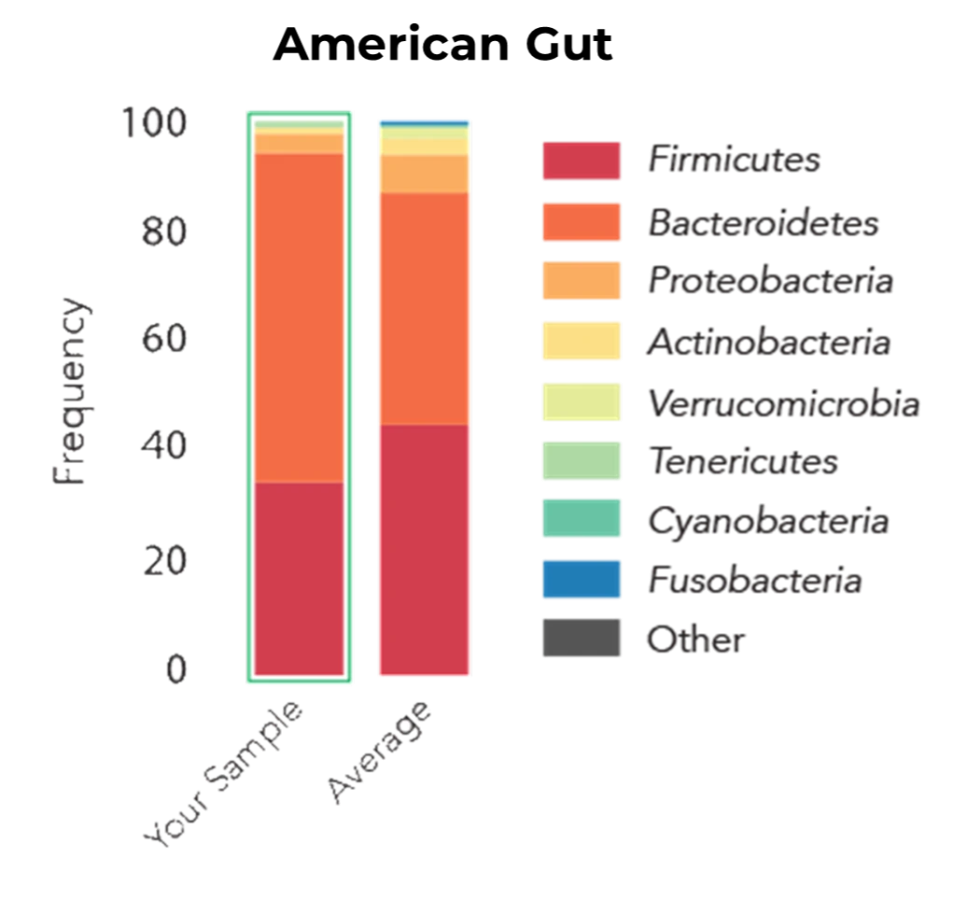
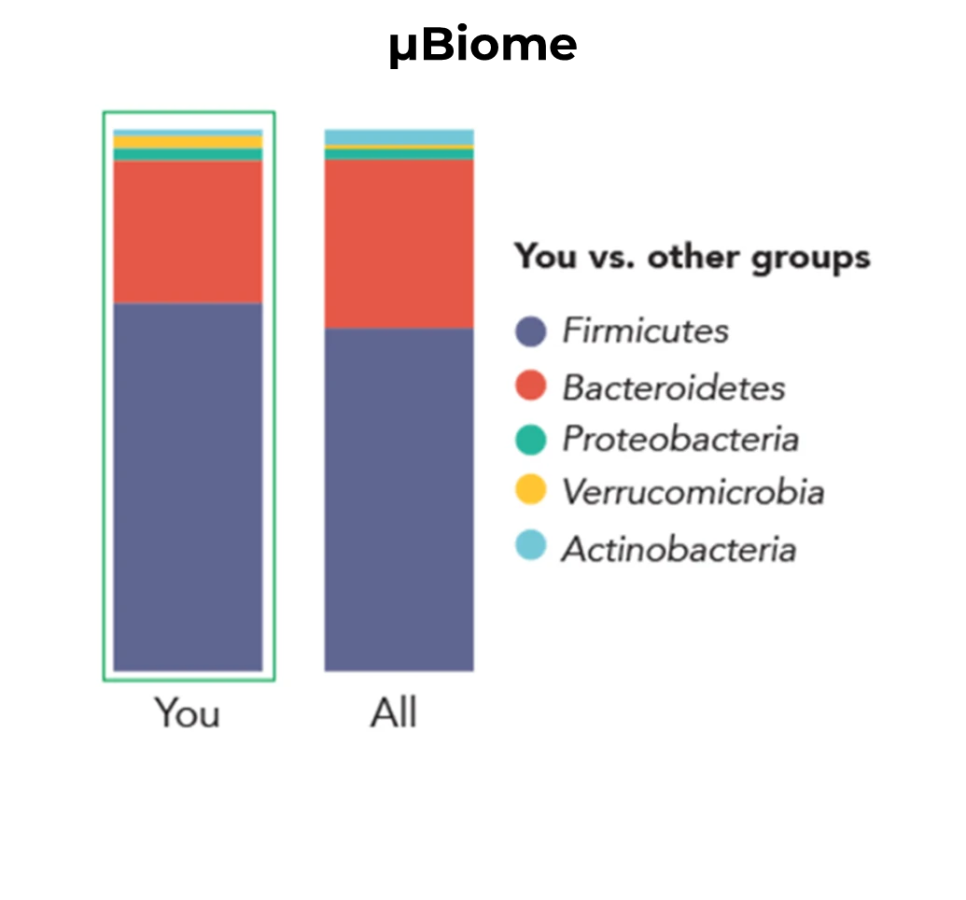
A real-world illustration of microbiome bias was described in a 2014 Science News article titled “Here’s the Poop on Getting Your Gut Microbiome Analysed.” In this case, the author submitted identical faecal samples to two different microbiome testing services and received dramatically different microbial profiles in return. The American Gut Project reported that Bacteroidetes, a group of Gram-negative bacteria, dominated both the sample. Meanwhile, results from µBiome suggested that Firmicutes, which are predominantly Gram-positive, were the most abundant group. One plausible interpretation of this discrepancy is that differences in lysis efficiency, particularly insufficient disruption of Gram-positive cells, led to an underrepresentation of Firmicutes in the American Gut analysis.
|
|
|
|---|
Identifying lysis issues before sequencing starts
Clues pointing to lysis-related problems can often be identified even before sequencing begins. For example, when processing a mock microbial community standard, an expected DNA yield might be 2 micrograms from 75 microliters of material. If the recovered DNA quantity is substantially lower, this deviation should prompt further investigation. Should sequencing data later reveal both a low yield and a skew toward Gram-negative organisms, this strongly suggests a failure to adequately lyse more resilient microbes. On the other hand, if DNA yield is reduced without any apparent taxonomic bias, then other variables such as DNA purification efficiency, recovery losses, or even inaccuracies in quantification methods should be evaluated.
The importance of routine standards and yield monitoring
To ensure consistency, it is best practice to process a standard reference sample routinely, ideally alongside every batch of experimental samples. Monitoring DNA yield across runs enables early detection of protocol changes or reagent issues. When yields suddenly drop under otherwise identical conditions and measurement techniques, it is a clear signal that something in the workflow has needs correction.
Amplification bias and GC content effects
Beyond DNA extraction, amplification bias represents another major contributor to variability in microbiome studies. Often referred to as GC bias, this issue stems from differences in genomic guanine-cytosine content across microbes. Because GC base pairs form stronger bonds than adenine-thymine pairs, DNA regions with high GC content are more difficult to denature during PCR. As a result, library preparation workflows with suboptimal PCR conditions, or those requiring excessive amplification, can preferentially distort representation. Although Firmicutes are generally characterised by lower GC content, making them easier to amplify, this same property could lead to their apparent overrepresentation under biased PCR conditions, as potentially observed in the µBiome results.
Why standardised controls are essential in microbiome research
Ultimately, without proper controls, it is impossible to determine precisely which biases are influencing a given dataset or how strongly they affect the final microbial profile. However, by analysing the same biological sample across different platforms, the author of the Science News article demonstrated why standardised controls are essential in microbiome research. Using a well-characterised reference standard rather than biologically variable samples such as faeces makes it possible to monitor bias, validate workflows, and correct issues before processing valuable experimental samples. Incorporating such standards is a critical step toward producing microbiome data that is both reliable and reproducible.
How Zymo Research Microbial Standards support accurate microbiome analysis
Zymo Research’s microbial standards are designed specifically to address the need for reliable, well-characterised reference materials in microbiome research. These standards consist of defined microbial compositions with known taxonomic profiles and controlled quantities, enabling researchers to objectively evaluate DNA extraction efficiency, amplification bias, and overall workflow performance. By incorporating organisms with a range of cell wall structures and lysis sensitivities, Zymo’s standards help reveal biases related to cell wall recalcitrance that might otherwise go unnoticed. When used routinely as internal controls, these reference materials allow researchers to benchmark protocols, track consistency over time, and identify sources of technical variation before they impact experimental samples. As a result, Zymo Research’s microbial standards provide a practical and effective foundation for improving data accuracy, reproducibility, and confidence in microbiome sequencing studies.
This content is inspired by Zymo's original blog about mastering the microbiome.