- In this section:
- Contact our Zymo custom service specialists
Cambridge Bioscience offers access to a site- or locus-specific 5-mC analysis service for epigenetic biomarker validation from Zymo Research. Providing publication quality data, the MethylCheck™ service is suitable whether you have methylation array (27K/450K/850K) data that you would like to validate in a large sample cohort or a specific gene region in mind. Zymo Research’s scientists are available to design, validate and evaluate site-specific DNA methylation changes. Simply send your samples and regions of interest (following discussion with Cambridge Bioscience) and Zymo Research will perform every step right through to data analysis.
Benefits of using the MethylCheck™ Service 
• Primer design and validation
• Targeted amplification
• Adapterisation and barcoding
• Illumina™ sequencing technology
• DNA methylation data bioinformatics analysis and support
Contact our specialist
To discuss your requirements or to find out more, please contact our Zymo Research specialist.
Cambridge Bioscience offers access to a range of single base resolution DNA methylation analysis services from Zymo Research. Offering a range of different coverage levels from genome wide to whole genome coverage, these services combine next-gen sequencing and innovative bioinformatics with well-established bisulfite technologies providing the most comprehensive DNA methylation analysis service available.
Benefits of using the single base resolution DNA methylation analysis services
• Next-generation bisulfite sequencing platforms for DNA methylation analysis
• Comprehensive bioinformatic analysis
• Low DNA input - less than 500ng genomic DNA
• Wide sample source compatibility
• Customisable service
• Rapid turnaround time
Services
The Methyl-MaxiSeq™, Methyl-MidiSeq™ and Methyl-MiniSeq™ services include:
• Sample processing and library preparation
• Next-generation sequencing
• Sequencing validation and bioinformatic analysis
• Data output and delivery
Methyl-Mini Seq™- covers ~10 percent of the methylome
The Methyl-MiniSeq™ platform (an improved version of Reduced Representation Bisulfite Sequencing for greater coverage) can be used to detect three to four million unique CpG sites, allowing over 85 percent coverage of all CpG islands and over 80 percent of all gene promoters for a maximal amount of methylation data from less sequencing reads, reducing the overall cost. The system is applicable to biomarker discovery by providing for the identification and analysis of differentially methylated regions (DMRs) between samples.
Methyl-Midi Seq™- covers ~30 percent of the methylome
Methyl-MidiSeq™ can be used to detect eight to nine million unique CpG sites. Extending the coverage of the Methyl-MiniSeq™ platform to include a large majority of genetic regulatory elements, gene bodies and repeated DNA sequences, it is a good option for those researchers requiring methylome analysis outside of gene promoters and CpG islands.
Methyl-MaxiSeq™- covers the entire methylome
The Methyl-MaxiSeq™ platform offers whole-genome bisulfite sequencing and is intended for the detection of DNA methylation across the entire genome. DNA methylation information is provided in CpG context as well as in the less common CHG and CHH contexts. Attaining an average read coverage of 15-20X per base (for the human genome), this can be modified depending on your requirements. Since whole-genome sequence is provided, SNP analysis can be also be performed simultaneously.
Genome-wide DNA methylation epigenetic analysis services comparison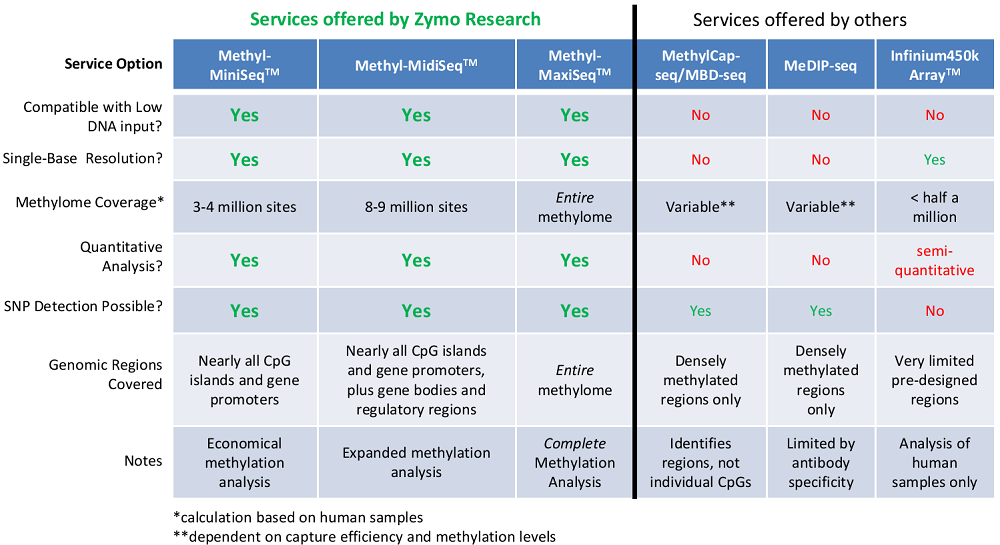
Speak to a specialist
For further information or for pricing for this service, please contact our Zymo custom services specialist.
Infinium is a registered trademark of Illumina.

